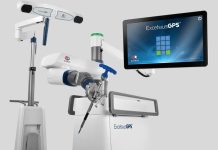
Revolutionary AI has upgraded the endeavor of protein identification into something akin to a quick-as-a-flash web search. Researchers are leveraging AlphaFold, a cutting edge AI network, to estimate the exact structures of more than 200 million proteins from around a million species. Given the thoroughness of this capability, nearly every protein on the planet can be accurately modeled in 3D for scientific study.
A new database assembled by London-based AI company DeepMind has made this massive data aggregate available for free. “Essentially you can think of it covering the entire protein universe,” said the company’s Chief Executive Officer, Demis Hassabis. “We’re at the beginning of a new era of digital biology.” The European Molecular Biology Laboratory’s European Bioinformatics Institute (EMBL–EBI), an intergovernmental organization in the U.K., also contributed to the construction of the database.
Cell functionality in proteins is based on the structure of an individual protein, and many groundbreaking drugs are derived from their structural metrics, which also imply how they function in life forms.
Deep learning techniques were integral to the development of the AlphaFold network, which has amassed approximately one million entries in its year of existence, up from a launch total of 350,000 that mostly consisted of proteins found in humans, mice, and around 20 other commonly studied organisms.